Correlation between Gene Expression and Histology in Transbronchial Lung Transplant Biopsies
University of Alberta, Edmonton, AB, Canada
Meeting: 2019 American Transplant Congress
Abstract number: C320
Keywords: Biopsy, Gene expression, Histology, Lung transplantation
Session Information
Session Name: Poster Session C: Lung: All Topics
Session Type: Poster Session
Date: Monday, June 3, 2019
Session Time: 6:00pm-7:00pm
 Presentation Time: 6:00pm-7:00pm
Presentation Time: 6:00pm-7:00pm
Location: Hall C & D
*Purpose: Transbronchial lung transplant biopsies (TBBx) are limited by non-specific histologic lesions and suboptimal interobserver reproducibility. Gene expression offers a potentially more precise, objective and mechanistic diagnostic approach, but it has not yet been fully utilized in lung transplants. The aim of this study was to assess the feasibility of gene expression in formalin-fixed paraffin-embedded (FFPE) lung transplant biopsies.
*Methods: All TBBx performed at a single institution between 01/2015 and 12/2017 were retrieved (n=43). Those without alveolar parenchyma (n=9) or residual FFPE tissue (n=2) were excluded. Concurrent mucosal biopsies (MBx) were also retrieved (n=11). Review diagnoses were assigned by a pathologist prior to acquisition of molecular data. NanoString was used to analyze a literature-derived 136-gene set, including previously-described allograft rejection and acute lung injury transcripts. Gene expression results were correlated with histology, biopsy characteristics and spirometry.
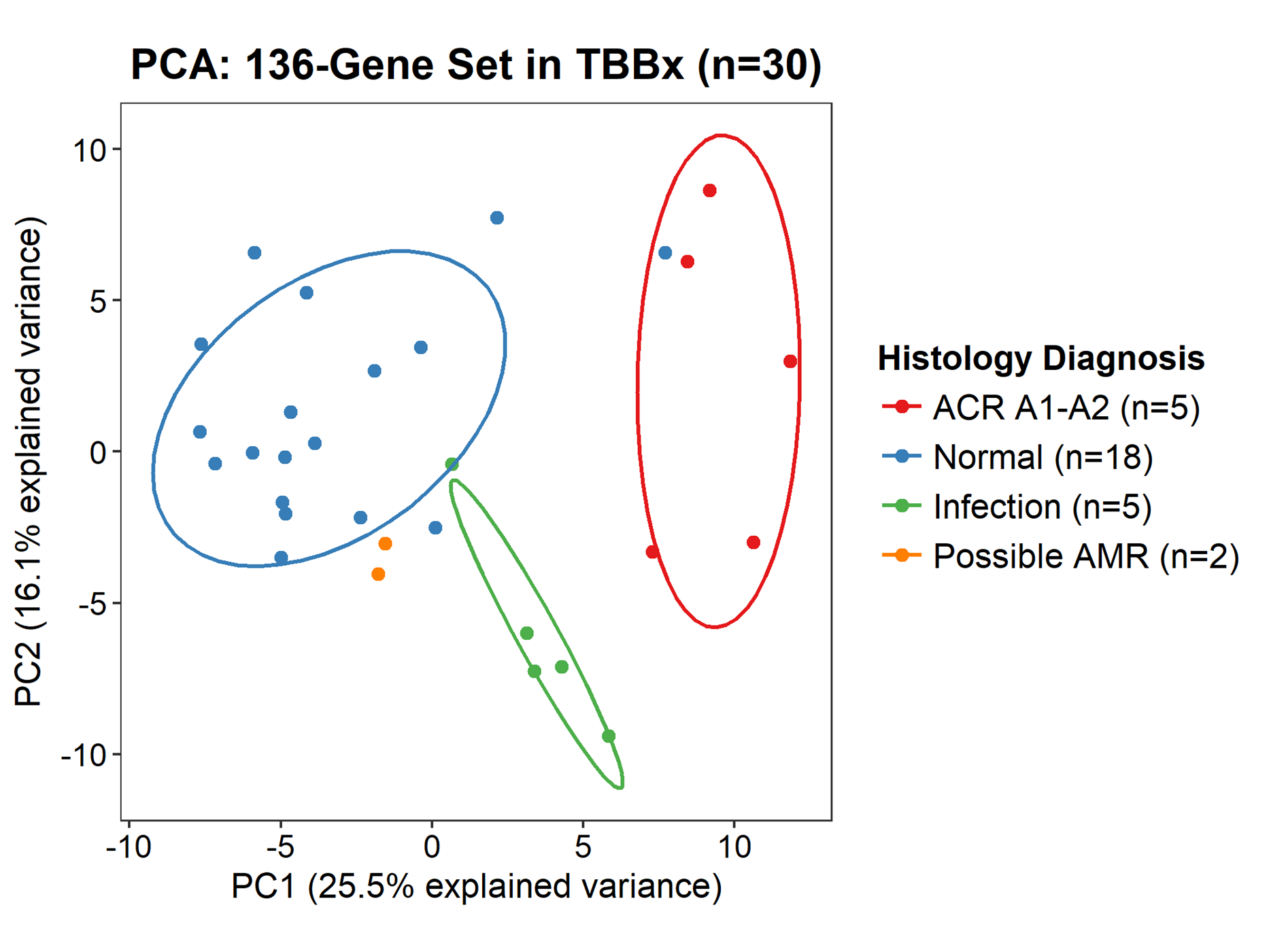
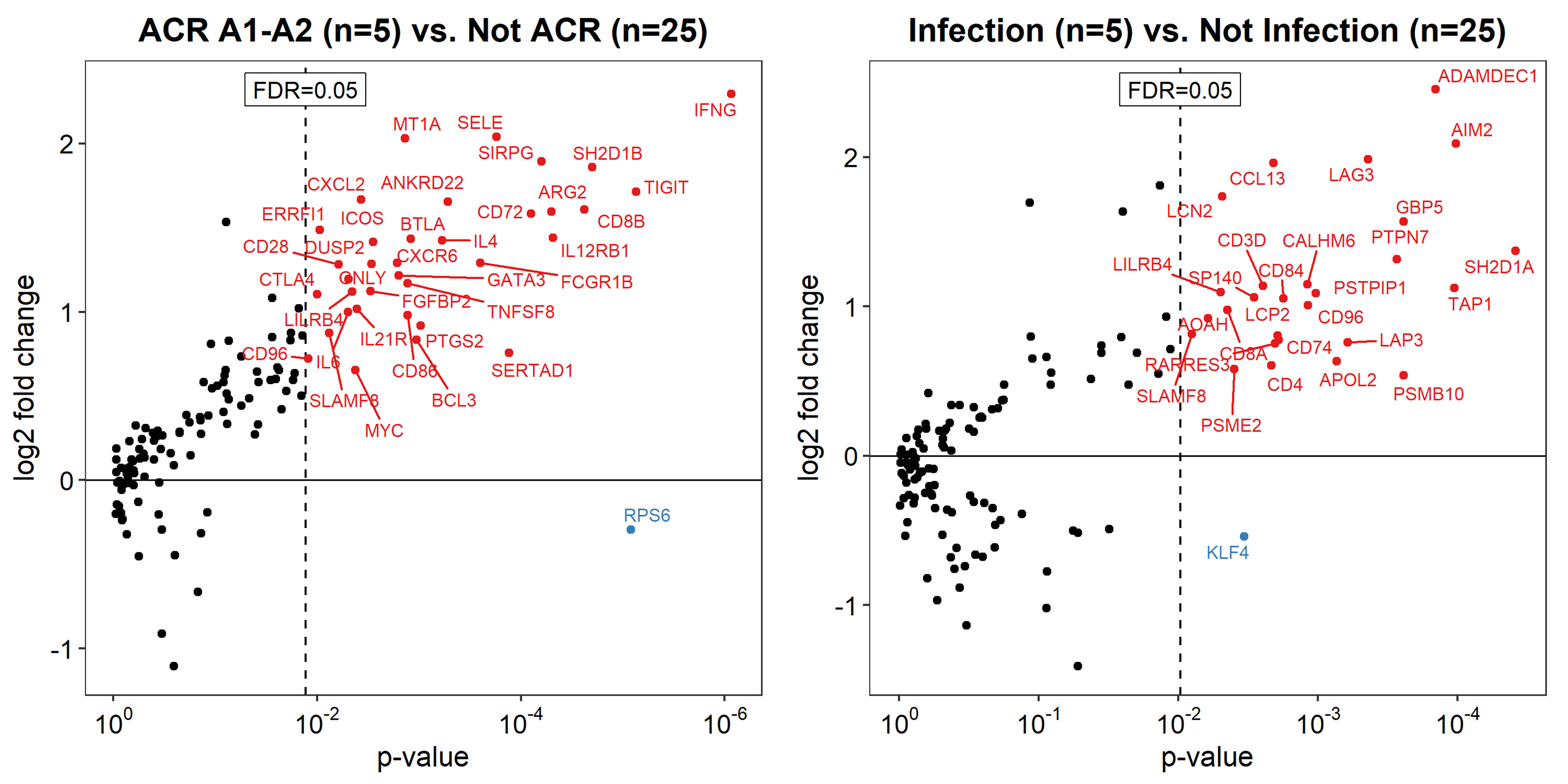
*Results: 30/32 TBBx and 11/11 MBx passed quality control and were included in the final analysis. TBBx were reclassified as acute cellular rejection (n=5), infection (n=5), possible antibody-mediated rejection (n=2) and normal (n=18). There were no significant differences in patient or biopsy characteristics between diagnostic groups (p>0.05). Unsupervised principal component analysis demonstrated a general correlation between gene expression patterns and histology diagnoses in TBBx (Figure 1) but not MBx. Surfactant gene expression correlated with the presence (p<0.001) and proportion (r=0.501, p=0.005) of alveolar parenchyma. Volcano plot analysis identified novel gene sets for acute cellular rejection and infection (FDR<0.05) (Figure 2). No correlation was identified between gene set expression and spirometry parameters (p>0.05).
*Conclusions: These results suggest that, for TBBx with alveolar parenchyma, gene expression patterns generally correlate with histologic categories. Molecular analysis of FFPE TBBx can potentially be used to identify and validate molecular signatures of clinical significance, refine histologic criteria, and provide ancillary diagnostic tools.
To cite this abstract in AMA style:
Adam B, Du K, Rotich S, Mengel M. Correlation between Gene Expression and Histology in Transbronchial Lung Transplant Biopsies [abstract]. Am J Transplant. 2019; 19 (suppl 3). https://atcmeetingabstracts.com/abstract/correlation-between-gene-expression-and-histology-in-transbronchial-lung-transplant-biopsies/. Accessed May 28, 2026.« Back to 2019 American Transplant Congress