Molecular Analysis of Acute Cellular Rejection in Clinical Face Transplants
Brigham and Women's Hospital, Boston.
Meeting: 2018 American Transplant Congress
Abstract number: 281
Keywords: Biopsy, Gene expression, Rejection, T cells
Session Information
Session Time: 2:30pm-4:00pm
 Presentation Time: 2:30pm-2:42pm
Presentation Time: 2:30pm-2:42pm
Location: Room 303
Although monitoring and immunosuppression of vascularized composite allografts (VCA) draw experience from solid organ transplantation, VCAs pose unique challenges. A deeper understanding of mechanisms underlying VCA rejection is critical for development of anti-rejection therapies targeted for VCA recipients. The availability of biopsies from the largest cohort of face transplant patients at a single center worldwide allowed us to study the molecular signature of acute cellular rejection (ACR).
NanoString platform was used to quantify the expression of 730 genes in skin biopsies collected from 7 face transplant patients during ACR (Banff grade 1, n=6; grade 2, n=8; grade 3, n=11) and non-rejection (Grade 0; n=11), and compared with facial skin biopsies from non-transplanted patients with rosacea (n=3) and delayed type hypersensitivity reaction (n=4), and healthy non-transplanted facial skin (n=4). Gene expression findings were validated at protein level using immunofluorescence staining of biopsies.
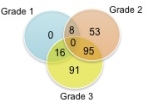
We found distinct gene expression profiles associated with ACR, inflammatory dermatoses and normal skin. Comparison of grade 3 ACR vs non-rejection biopsies identified 202 differentially expressed genes (adjusted p value<0.05). Functional pathway analysis showed that 3 gene sets are over-expressed in ACR: T cell activation, interferon-gamma responses, and cytotoxicity. We identified distinct gene expression signatures in biopsies that correlated significantly with each of the different grades of rejection (Fig1). Analysis of unique genes revealed 142 genes that are differentially expressed exclusively in face transplants during rejection, but not in inflammatory dermatoses or normal skin.
This is the most comprehensive study to date to identify the molecular signature of ACR in clinical VCAs. Our findings suggest that skin biopsies from face transplants during ACR, although indistinguishable on histologic analysis from inflammatory dermatoses, reveal extensive differences at the molecular level.
Fig1. Differences in gene expression progress steadily with worsening grades of rejection. All comparisons made versus Grade 0.
CITATION INFORMATION: Win T., Dyring-Andersen B., Teague J., Murakami N., Chandraker A., Pomahac B., Riella L., Clark R. Molecular Analysis of Acute Cellular Rejection in Clinical Face Transplants Am J Transplant. 2017;17 (suppl 3).
To cite this abstract in AMA style:
Win T, Dyring-Andersen B, Teague J, Murakami N, Chandraker A, Pomahac B, Riella L, Clark R. Molecular Analysis of Acute Cellular Rejection in Clinical Face Transplants [abstract]. https://atcmeetingabstracts.com/abstract/molecular-analysis-of-acute-cellular-rejection-in-clinical-face-transplants/. Accessed May 19, 2026.« Back to 2018 American Transplant Congress