The Microenvironment of Belatacept-Resistant Rejection
Surgery, Emory University, Atlanta, GA
Meeting: 2021 American Transplant Congress
Abstract number: 504
Keywords: Co-stimulation, Image analysis, Rejection, T cell receptors (TcR)
Topic: Basic Science » Acute Rejection
Session Information
Session Name: Acute Rejection
Session Type: Poster Abstract
Session Date & Time: None. Available on demand.
Location: Virtual
*Purpose: Belatacept is a co-stimulation blockade immunosuppressant, that despite demonstrating improved long-term outcomes, has not been widely adopted, in part, due to concern over early episodes of acute rejection. A method of co-immunosuppression with transient calcineurin inhibitor (CNI) therapy has improved rejection rates, but importantly, has also clearly defined a population of patients well controlled on CNIs but who develop rejection when that therapy is withdrawn. We aim to characterize the microenvironment of that rejection.
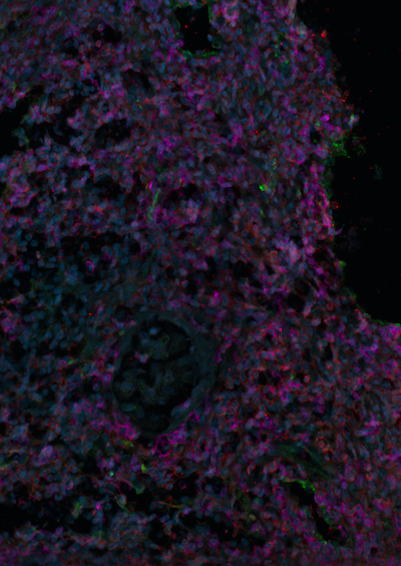
*Methods: We selected formalin fixed paraffin embedded biopsies from 5 historical patients with Banff 2B acute rejection in the setting of CNI weaning. From these biopsies, we sequenced the β chain of T cell receptors (TCR) to analyze clonality of the T cells infiltrating biopsies. TCR sequences were analyzed with the R biostatistical package immunarch as well as Adaptive Biotechnologies ImmunoSeq Analyzer tool. In addition, we performed multiplex immunofluorescence to characterize the immune cells infiltrating these biopsies. Primary antibodies against CD4, CD8 and MHCII were used along with a DAPI stain to allow cell identification. Slides were imaged with a Nikon Leica Spinning Disk microscope, and image processing performed using Fiji software.
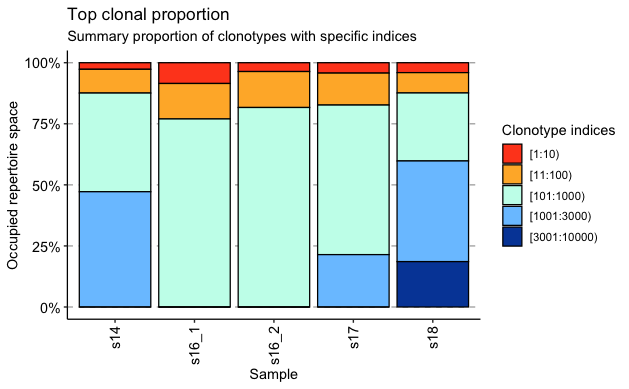
*Results: There was an average of 3078 total and 2542 productive templates per sample, with 82% of templates productive. Productive Simpson clonality ranged from 0.02 to 0.057. The maximum productive frequency in sample s16_1 was 4.5%. The remaining samples had maximum productive frequencies between 0.4 and 1.1%. There was minimal repertoire overlap between samples. Analysis of V and J gene usage revealed a high frequency of BJ02-07 usage in all samples. Immunofluorescence imaging revealed a high density of both CD4 and CD8 cells, with a paucity of MHCII expressing cells.
*Conclusions: Though individual clones do not predominate in these biopsies, V/J gene usage suggests high levels of similarity among clones. We plan to quantify this with CDRdist scoring, a method based on Smith-Waterman alignment calculation. Qualitative immunofluorescence data will be processed with CellProfiler software to analyze spatial relationships between cell types.
To cite this abstract in AMA style:
Johnson A, Zhang J, Larsen C. The Microenvironment of Belatacept-Resistant Rejection [abstract]. Am J Transplant. 2021; 21 (suppl 3). https://atcmeetingabstracts.com/abstract/the-microenvironment-of-belatacept-resistant-rejection/. Accessed May 19, 2026.« Back to 2021 American Transplant Congress