The Metagenomic Landscape of Renal Transplant
A. Johnson1, G. Karadkhele2, C. Larsen2
1Emory University, Atlanta, GA, 2Surgery, Emory University, Atlanta, GA
Meeting: 2021 American Transplant Congress
Abstract number: 164
Keywords: Genomics, Immunosuppression, Infection
Topic: Clinical Science » Infectious Disease » Kidney Infectious Non-Polyoma & Non-Viral Hepatitis
Session Information
Session Name: Infections in Kidney Recipients
Session Type: Rapid Fire Oral Abstract
Date: Sunday, June 6, 2021
Session Time: 6:00pm-7:00pm
 Presentation Time: 6:30pm-6:35pm
Presentation Time: 6:30pm-6:35pm
Location: Virtual
*Purpose: The selective targeting of T cells in transplant immunosuppression medications leads to a high rate of viral infections. Different immunosuppression regimens have been shown to have variable effects on protective immunity. We aim to use next-generation sequencing (NGS) to characterize the microvirome of renal transplant patients and examine variance in the context of immunosuppression, rejection, and clinically apparent infection.
*Methods: Plasma samples are collected at baseline and at regular intervals after kidney transplant. Three patients with a total of 21 samples were selected as a pilot cohort. These patients were selected because of known clinically significant and quantified viremias (BK and CMV). Cell-free DNA was isolated from plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen). The isolated DNA was fragmented and appended with dual-indexed bar codes using the NexteraXT DNA Library Preparation kit (Illumina). Libraries were validated by capillary electrophoresis on an Agilent 4200 TapeStation, pooled at equimolar concentrations, and sequenced on an Illumina NovaSeq 6000 at 150PE to a depth of approximately 30 million reads/sample. Low quality reads were computationally removed, and adapters trimmed. Host and contaminant reads were removed, and the remaining sequences aligned against the known virome.
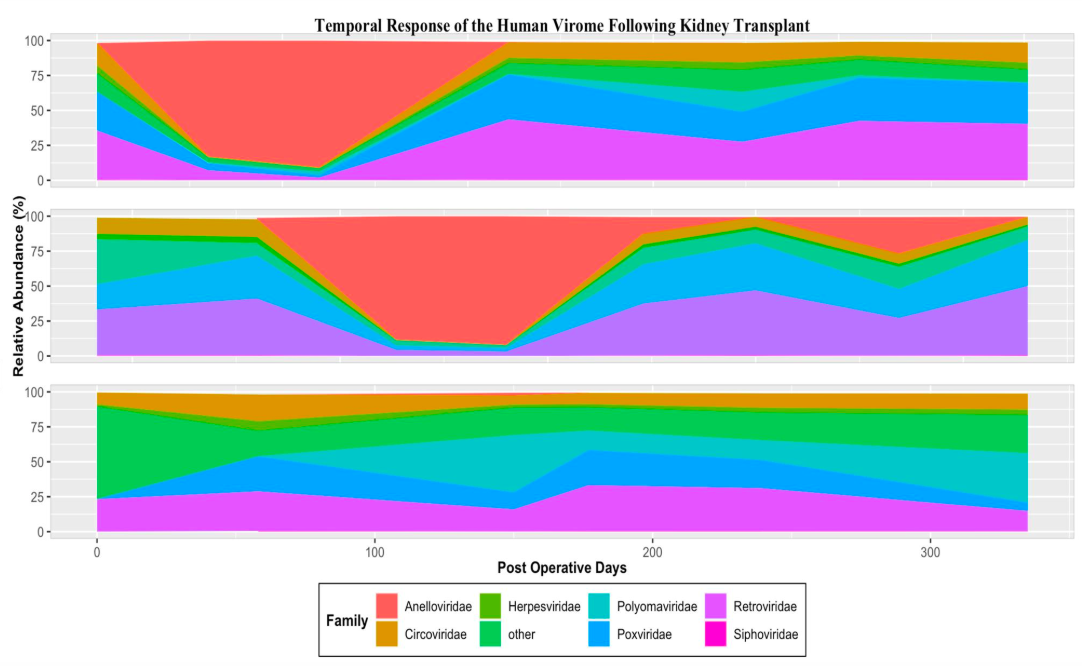
*Results: After removal of human, contaminant, and low-quality reads, an average of approximately 400,000 reads per sample remained, representing only 2% of overall reads. Total counts of BK virus or CMV obtained from NGS were compared to viral load levels obtained clinically. The two measurements demonstrated parallel trajectories, confirming the ability of NGS methods to detect and quantify clinically significant viremias. Aligned sequences were plotted by relative abundance over time for each sample. Two samples demonstrated notable expansion of anelloviridae during the first year after transplant. In addition to anelloviridae, herpesviridae, and polyomaviridae, samples also had significant levels of retroviridae and poxviridae.
*Conclusions: The microvirome is an underutilized tool to understand protective immunity in the transplant population. We plan to expand this method to banked samples for an additional 20 kidney transplant patients to study the influence of medications and other clinical features on the microvirome. We will also use a complementary contig assembly method to strengthen the mapping of reads to the known virome.
To cite this abstract in AMA style:
Johnson A, Karadkhele G, Larsen C. The Metagenomic Landscape of Renal Transplant [abstract]. Am J Transplant. 2021; 21 (suppl 3). https://atcmeetingabstracts.com/abstract/the-metagenomic-landscape-of-renal-transplant/. Accessed May 25, 2026.« Back to 2021 American Transplant Congress