Single-Cell RNA-seq Identifies Intra-Graft Population Heterogeneity in Acute Heart Allograft Rejection in the Mouse
1The First Affiliated Hospital,Sun Yat-sen University, Guangzhou, China, 2Immunobiology & Transplant Sciences Houston Methodist Hospital, Houston, TX
Meeting: 2021 American Transplant Congress
Abstract number: 443
Keywords: Endothelial cells, Mice, Rejection
Topic: Basic Science » Acute Rejection
Session Information
Session Time: 7:30pm-8:30pm
 Presentation Time: 7:40pm-7:50pm
Presentation Time: 7:40pm-7:50pm
Location: Virtual
*Purpose: Transplant rejection remains a major barrier to graft survival and involves a diversity of cell types. But the heterogeneity in a given cell type in graft rejection remains poorly defined.
*Methods: In the present study, we used single-cell RNA sequencing (scRNA-seq) technology, which is a powerful tool in identifying population heterogeneity, to examine changes in graft infiltrating cells during acute heart allograft rejection.
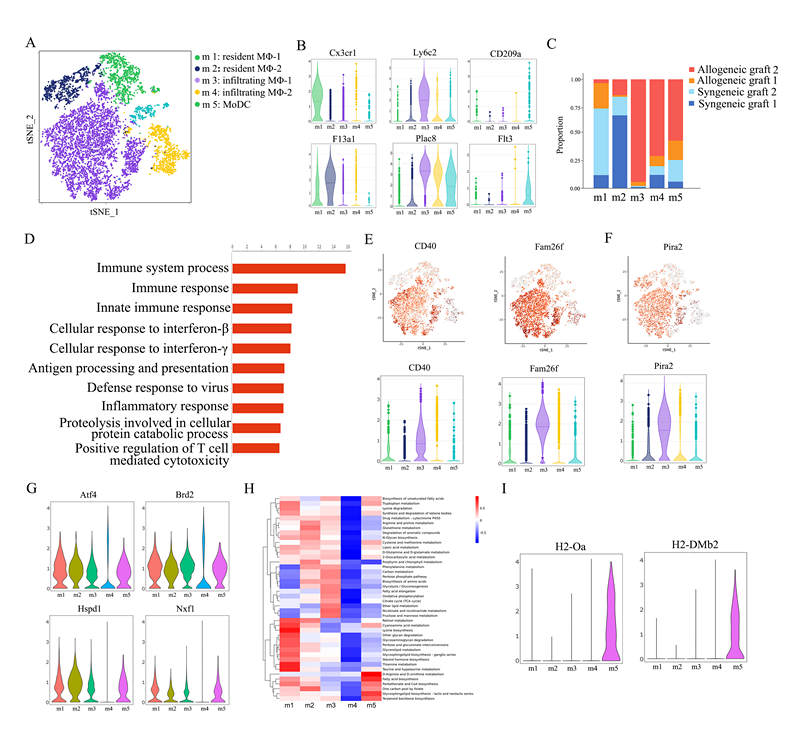
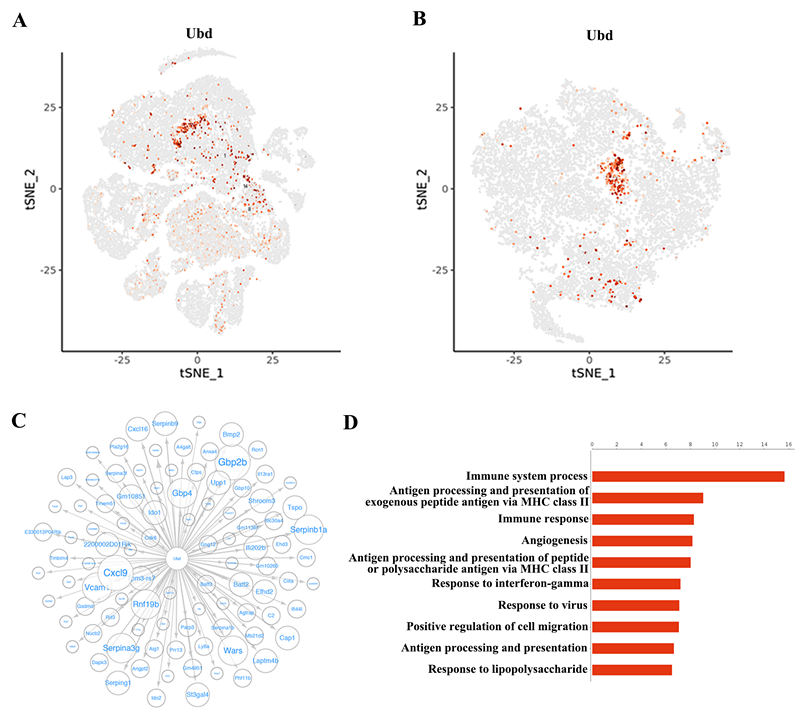
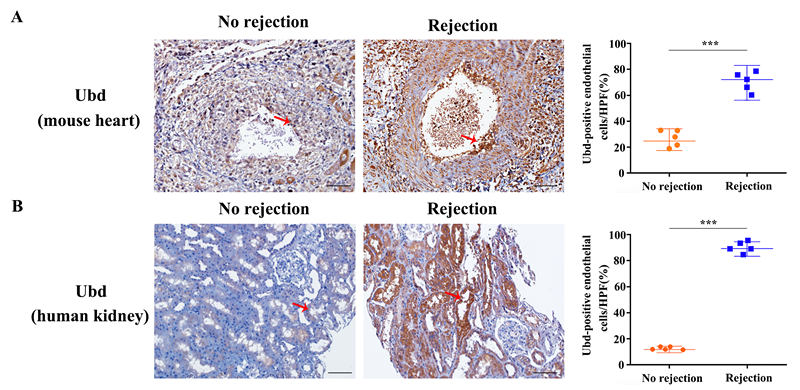
*Results: Here we focused on innate cells and performed the first comprehensive analysis of transcriptional heterogeneity in the mouse. In dimensionality reduction and unsupervised cell clustering analysis, we identified 21 distinct cell populations in the allografts. We showed that macrophages can be divided into five subclusters and revealed a new subset of macrophages highly associated with graft rejection; they upregulated CD40, Fam26f and Pira2. Furthermore, we confirmed the heterogeneity of graft endothelial cells in rejection, which showed five subclusters (EC1~EC5) and the EC5 can function as antigen presenting cells during transplant rejection. We further identified Ubiquitin D (Ubd) in EC5 cluster as a key marker of rejection, which was validated by immunohistochemistry staining in mouse cardiac samples and human kidney biopsy specimens.
*Conclusions: our findings revealed the population heterogeneity of cell types in acute allograft rejection and identified novel markers in rejection.
To cite this abstract in AMA style:
Wu C, Tang Y, Shi X, He X, Li X. Single-Cell RNA-seq Identifies Intra-Graft Population Heterogeneity in Acute Heart Allograft Rejection in the Mouse [abstract]. Am J Transplant. 2021; 21 (suppl 3). https://atcmeetingabstracts.com/abstract/single-cell-rna-seq-identifies-intra-graft-population-heterogeneity-in-acute-heart-allograft-rejection-in-the-mouse/. Accessed May 18, 2026.« Back to 2021 American Transplant Congress