Gene Expression Levels of Advanced Heart Failure Patients in Terms of Coagulation and Its Effect within Multiple Organ Dysfunction Syndrome.
Medicine Cardiology, UCLA, Los Angeles, CA.
Meeting: 2016 American Transplant Congress
Abstract number: B144
Keywords: Anticoagulation, Gene expression, Heart failure, Outcome
Session Information
Session Name: Poster Session B: Hearts and VADs in Depth - The Force Awakens
Session Type: Poster Session
Date: Sunday, June 12, 2016
Session Time: 6:00pm-7:00pm
 Presentation Time: 6:00pm-7:00pm
Presentation Time: 6:00pm-7:00pm
Location: Halls C&D
MOD is progressive organ failure amongst 2 or more organs. Innate-immunity/inflammation-related coagulopathies can cause disseminated clotting in vasculature leading to organ failure. Purpose is to depict MOD development and identify biomarkers to improve clinical diagnosis and allocate resources effectively. We hypothesize differentially expressed genes within coagulation module in high and low risk MCS patients against controls. 22 MCS implanted patients and 4 healthy CTRLs were enrolled in study. Peripheral Blood Mononuclear Cells (PBMC) collected relative to surgery with time points -1, +1, +3, +5, +8 days later were subjected to whole genome mRNA sequencing. Weighed Gene Co-expression Network Analysis (WGCNA) was used to identify module-associated eigengenes. Two-Way ANOVA with Benjamini-Hochberg correction and Fold Change were used to determine significant genes amongst INTERMACS1&2(High), INTERMACS3&4(Low), and CTRL Groups. WGCNA identified 20 eigengene modules. Statistical analyses applied to 2 modules (3117 genes) and resulted 380 coagulation-related genes (q<0.05). 230 genes exhibited 2.0 fold change. 26 inflammation/coagulation-related genes have significant enrichment in 271 GO categories (q-value <0.05): regulation of body fluid levels, myeloid leukocyte migration, leukocyte cell-cell adhesion, regulation of blood vessel morphogenesis and endothelial cell migration. High risk group displayed up-regulation of PROS1, ITGB3, TREML1, CLU, GP6; down-regulation of LAT,T-cell antigen receptor, and of PDGFB, platelet-derived growth factor receptor for pericyte recruitment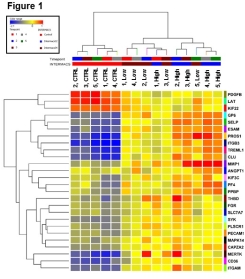 . Endothelial cells (SELP and ESAM) play essential role in coagulopathy through regulation of binding sites amongst anticoagulant and pro-coagulant factors on cell surface. Understanding coagulation may play pivotal role in outcome prediction following MCS implant. Using bioinformatics tools, differentially expressed genes within AdHF patients in comparison to CTRL could show promising leads in better comprehending pathophysiology of MOD. Further analysis necessary for validation.
. Endothelial cells (SELP and ESAM) play essential role in coagulopathy through regulation of binding sites amongst anticoagulant and pro-coagulant factors on cell surface. Understanding coagulation may play pivotal role in outcome prediction following MCS implant. Using bioinformatics tools, differentially expressed genes within AdHF patients in comparison to CTRL could show promising leads in better comprehending pathophysiology of MOD. Further analysis necessary for validation.
CITATION INFORMATION: Togashi R, Chang E, Wisniewski N, Cadeiras M, Bondar G, Reed E, Deng M. Gene Expression Levels of Advanced Heart Failure Patients in Terms of Coagulation and Its Effect within Multiple Organ Dysfunction Syndrome. Am J Transplant. 2016;16 (suppl 3).
To cite this abstract in AMA style:
Togashi R, Chang E, Wisniewski N, Cadeiras M, Bondar G, Reed E, Deng M. Gene Expression Levels of Advanced Heart Failure Patients in Terms of Coagulation and Its Effect within Multiple Organ Dysfunction Syndrome. [abstract]. Am J Transplant. 2016; 16 (suppl 3). https://atcmeetingabstracts.com/abstract/gene-expression-levels-of-advanced-heart-failure-patients-in-terms-of-coagulation-and-its-effect-within-multiple-organ-dysfunction-syndrome/. Accessed May 19, 2026.« Back to 2016 American Transplant Congress
