DSA Causes Mild Molecular ABMR-like Changes in Many Biopsies Not Diagnosed as Rejection
1Alberta Transplant Applied Genomics Centre, Edmonton, AB, Canada, 2., ., AB, Canada
Meeting: 2021 American Transplant Congress
Abstract number: 317
Keywords: Alloantibodies, Kidney, Kidney transplantation, Rejection
Topic: Clinical Science » Kidney » Kidney Chronic Antibody Mediated Rejection
Session Information
Session Name: Kidney Antibody Mediated Rejection
Session Type: Rapid Fire Oral Abstract
Date: Tuesday, June 8, 2021
Session Time: 4:30pm-5:30pm
 Presentation Time: 4:30pm-4:35pm
Presentation Time: 4:30pm-4:35pm
Location: Virtual
*Purpose: In the multicenter study INTERCOMEX, we developed a microarray-based system (MMDx) for diagnosing antibody-mediated and T cell-mediated rejection (ABMR, TCMR) in kidney transplant biopsies using an ensemble of machine-learning classifiers. Here we use an expanded set of 1679 biopsies to explore the significance of DSA positivity in kidneys without rejection.
*Methods: Machine learning was used as previously described, based on gene expression (Am J Transplant. 2019;19:2719). A new DSA classifier (DSAprob) was trained on DSA+ vs DSA- samples, with probabilities assigned in the left-out folds of 10-fold cross-validation. We plotted the biopsy samples’ molecular distribution using Uniform Manifold Approximation and Projection (UMAP), a recently developed dimensionality reduction method. The inputs were seven molecular rejection scores, built for predicting TCMR, ABMR, and i-, t-, ptc-, g-, and cg-lesions.
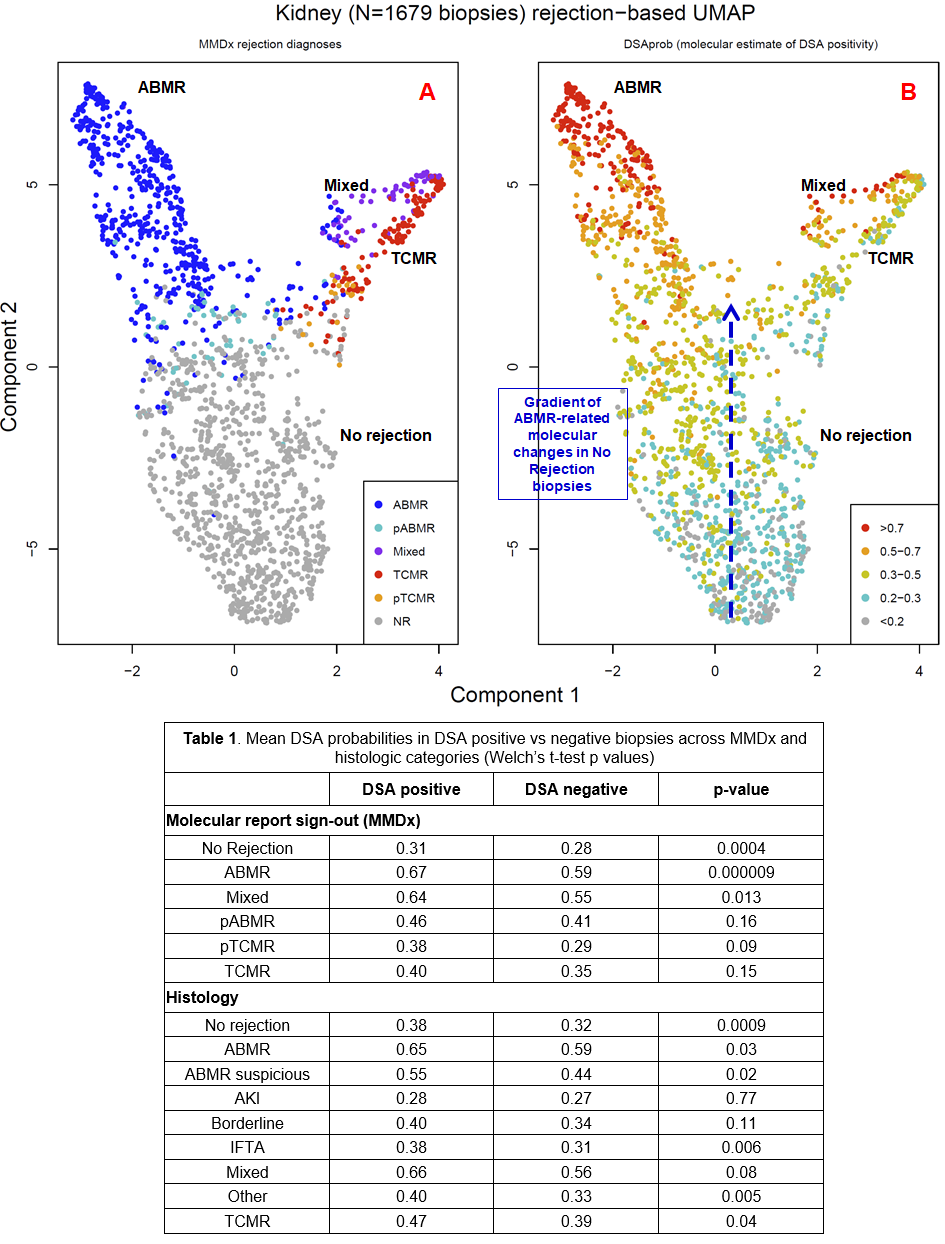
*Results: Figure 1A is a UMAP plot colored by MMDx diagnoses, mapping ABMR, TCMR, possible rejection (pABMR, pTCMR), Mixed, and No rejection. Figure 1B is the same distribution colored by DSAprob classifier scores, showing that a gradient of mild ABMR-related changes exists within No Rejection biopsies. There was also a gradient within ABMR correlating with stage (early>late) and in TCMR associated with Mixed rejection.
Within ABMR, DSAprob was higher in DSA+ than DSA- biopsies, but mild DSA-related changes were also detected in biopsies with No Rejection (NR), either by MMDx or histology (Table 1). DSAprob predicted DSA status, indicating the molecular changes detected in NR biopsies were DSA-related. In 738 biopsies with no rejection by MMDx, the 369 with high DSAprob scores (above the median) were more often DSA positive (36%) than those with low DSAprob scores (24%) (chisq pval=0.0004). Thus ABMR-related gene expression changes in No rejection biopsies are associated with DSA positivity.
NR biopsies with high DSAprob scores also had increased scores for other ABMR and NK-related classifiers and gene sets, but not TCMR or injury-related classifiers and gene sets.
*Conclusions: A gradient of mild ABMR-related gene expression including NK-related and IFNG-related changes exists in many kidneys currently called no rejection that corresponds to a higher frequency of DSA, indicating that mild molecular ABMR-related changes occur in many more kidneys than currently diagnosed as ABMR by MMDx or by histology. DSA is likely stressing the microcirculation more widely than previously realized. (ClinicalTrials.gov NCT01299168 (kidney).
To cite this abstract in AMA style:
Halloran PF, Madill-Thomsen KS. DSA Causes Mild Molecular ABMR-like Changes in Many Biopsies Not Diagnosed as Rejection [abstract]. Am J Transplant. 2021; 21 (suppl 3). https://atcmeetingabstracts.com/abstract/dsa-causes-mild-molecular-abmr-like-changes-in-many-biopsies-not-diagnosed-as-rejection/. Accessed May 31, 2026.« Back to 2021 American Transplant Congress